Origin and genomic evolution of polyploid complexes in Melampodium (Asteraceae)
A combination of methods and techniques is used to analyze in detail the origin and evolution of allotetraploids and allohexaploids of Melampodium. Phylogenetic and phylogeographic approaches are employed to elucidate the evolutionary origin and history of this complex, and are complemented by in-depth analyses of genome evolution following polyploidization, with a focus on repetitive DNA. The comparative investigation of this polyploid complex addresses evolutionary questions concerning the impact of origin and evolutionary history of polyploids on the type and frequency of genomic changes contributing to speciation.
Publications
McCann J, Macas J, Novak P, Stuessy TF, Villasenor JL, Weiss-Schneeweiss H. (2020) Differential genome size and repetitive DNA evolution in diploid species of Melampodium sect. Melampodium (Asteraceae). Frontiers of Plant Science 11:362.
McCann J, Jang T-S, Macas J, Schneeweiss GM, Matzke NJ, Novak P, Stuessy TF, Villasenor JL, Weiss-Schneeweiss H (2018) Dating the species network: Allopolyploidy and repetitive DNA evolution in American daisies (Melampodium sect. Melampodium, Asteraceae). Systematic Biology 67: 1010-1024.
McCann J, Schneeweiss GM, Stuessy TF, Villaseñor JL, Weiss-Schneeweiss H (2016) The impact of reconstruction methods, phylogenetic uncertainty and branch lengths on inference of chromosome number evolution in American daisies (Melampodium, Asteraceae). PLoS ONE 11: e0162299.
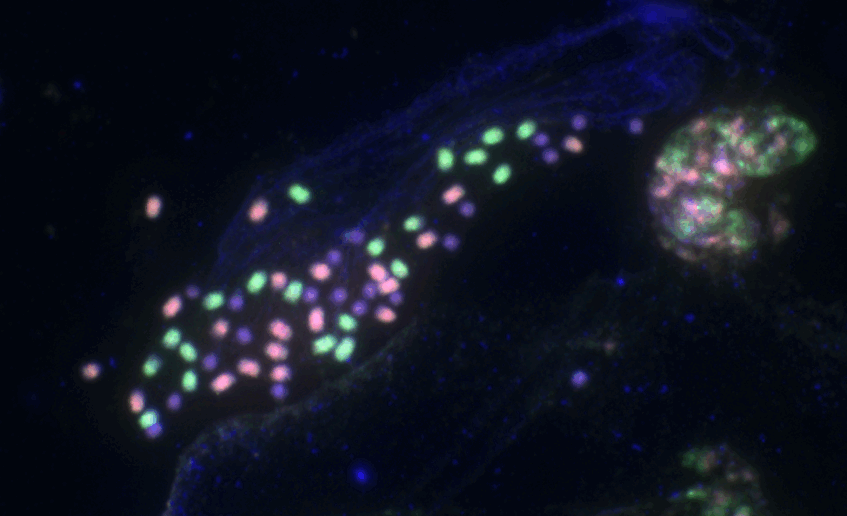
Jang T-S, Weiss-Schneeweiss H (2015) Formamide-free genomic in situ hybridization (ff-GISH) allows unambiguous discrimination of highly similar parental genomes in diploid hybrids and allopolyploids. Cytogenetic and Genome Research 146: 325–331.
Weiss-Schneeweiss H, Emadzade K, Jang T-S, Schneeweiss GM (2013) Evolutionary consequences, constraints, and potential of polyploidy in plants. Themed Issue of Cytogenetics and Genome Research “Trends in polyploidy research in animals and plants” 140: 137–150.
Weiss-Schneeweiss H, Blöch C, Turner B, Villaseñor JL, Stuessy TF, Schneeweiss GM (2012) The promiscuous and the chaste: frequent allopolyploid speciation and its genomic consequences in American daisies (Melampodium sect. Melampodium; Asteraceae). Evolution 66-1: 211–228.
